Quick look at Psomagen GutBiome+
After trying Psomagen’s low-cost 16S microbiome test I next tried their whole genome sequencing (WGS) version called GutBiome+. Priced only slightly higher than the 16S test ($129 vs. $99 on Amazon), GutBiome+ is a full metagenomic test that can see everything in your gut, down to the species and even strain level. The cheaper 16S test can’t reliably detect species and is generally only useful for a quick snapshot of your overall microbiome, but this one should be able to see much detail.
The basic test collection is the same: you do your business and send a tiny swab back to the lab using the included prepaid envelope. My report arrived exactly three weeks later (and 10 days faster than a kit I sent to Viome the same day using the same sample).
Overall Results
The GutBiome+ report is generated using a bioinformatics pipeline Psomagen licensed from Microba, the well-regarded Australian microbiome sequencing company, but the report itself is quite different. In general, the Psomagen report tries to give more actionable information than Microba, mostly around dietary recommendations, whereas Microba seems to rely more on their human coaches to help you decide what to do.
Psomagen summarizes my results into a single “Gut Health Score”, a weighted combination of several more specific variables, including their interpretation of some of my responses to their questionnaire.
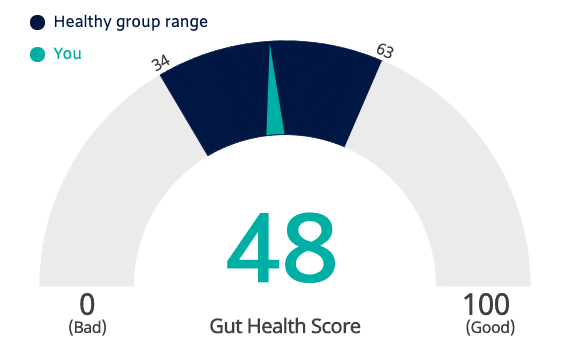
At first glance I was disappointed to see a number so far from 100, but the company explains that the “Healthy” comparison range goes from 34 to 63. Paradoxically I guess this means that anything over 63 is suspect, though the report itself isn’t clear about that.
Based on my overall score, Psomagen recommends a few food ideas to promote the health of my microbiome:

Interestingly, every single one of these is a high FODMAP food that caused great distress on my gut before I switched to the low-FODMAP diet I credit with my quick and complete recovery.
The other components of the Gut Health Score include Lifestyle Status (57), Nutrition Utility (74), Metabolic Supply (56), Metabolic Concern (38), and Microbiome Profile (80).

Lifestyle Status
Psomagen found only one area that would improve my overall sense of energy and lifestyle status. My D-Lactic Acid Production (25%) is slightly higher than the 22% that their healthy range predicts. Psomagen thinks I’ll have less fatigue if I improve that number. They suggest I eat smaller food portions throughout the day — which again, goes against the SIBO advice which suggests you that frequent snacking only promotes bloating by giving the out-of-place microbes too much to eat.
Nutrition Utility and Metabolic Supply
This score measures how well my microbiome breaks down food into the metabolites that drive health. Microbes play a major role in breaking down protein, starch, and fiber, and the report says that all my numbers seem to be doing well.
The rest of my metabolism — how my microbiome produces necessary vitamins and other digestive factors like GABA and Propionate — all seems fine according to the report. Considering that I took this test while nearly doubled over in pain from bloating and distention, the idea that all is well is hardly reassuring.

Metabolic Concern
At last I reach something that indicates a problem. My score, 38, is below the healthy range (41-69). Confusingly, according to the report “a high score means beneficial/non-beneficial metabolites balance is unstable by dysbiosis.” Hmmm, I’m not sure what that means, and anyway I have a low score. Does that mean my situation needs improvement, or am I in better shape than average? It’s unclear, but being out of the healthy range doesn’t look good.
Sure enough, diving into the details Psomagen points to several variables that indeed look like what I’d expect from a gut that was at the time suffering from distention and bloating. My ammonia production was higher than their healthy group average, and my Lipopolysaccharide production and protein degradation numbers were off-the-charts higher. Similarly, my TMAO production was much too high as well, as was my consumption of GABA.
Although it wasn’t clear that Psomagen’s advice was correct, I take the out-of-range results here to mean that something is out of whack.
### Microbe Profile
The report summarizes my overall microbiome diversity (80, which apparently is within the healthy range)
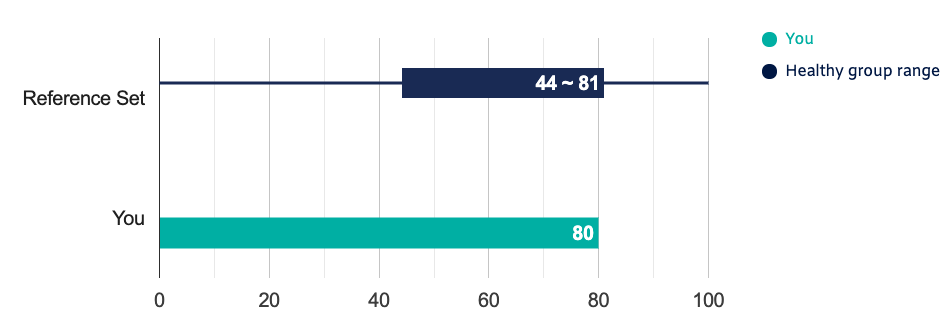
There is also a brief breakdown of the percentage of genes that are from bacteria, archaea, human, and novel. My phylum-level percentages roughly match what I’ve seen in other test results I’ve done over the years. The gut microbiome taxonomy is remarkably robust, despite my tummy feeling worse than it’s felt in years:
| Phylum | % |
|---|---|
| Firmicutes | 49.7 |
| Bacteroidetes | 14.4 |
| Actinobactera | 1.1 |
| Proteobacteria | 5 |
Apparently a full 24% of my gut is made of microbes that are not in the database; this is not an unusual WGS result.
I found it frustrating to scroll through the messy web-based UI to find the names and abundances of specific microbes, so I ended up doing a simple screen-scraping on the page HTML and made an Excel sheet that summarized the various microbes found. From that I was able to deduce that Of the 112 individual species they identified, only one was low out-of-range: Faecalibacterium sp (I was 0.1, whereas their range goes from 0.09 to 1.3) About half were “High”.
The most interesting result I found was very high levels of the proteobacteria Leclercia adecarboxylata. A 2014 review says it may be an opportunistic pathogen. Another article claims is may carry antibiotic resistant genes, and its unusually high abundance makes me wonder if I’m seeing some aftereffects from my surgery. Definitely a microbe to watch for in my next results. Incidentally it didn’t seem to be present in my previous Microba test, nor was there anything like it in my Viome results.
Summary
Psomagen’s metagenomic test is a good value at $120, but for most consumers a test is only as good as the report, and here’s where unfortunately the product currently falls short. Despite my clear digestive issues at the time I took the sample, the report advises dietary choices that were nearly opposite of what I should have been eating. Only by digging into the details was I able to find specific problems – seriously out-of-range metabolites – but it was difficult to tie those to specific microbes that may have offered treatment clues.
Although they’ll give you the raw FASTQ results if you request them, Psomagen’s reliance on the Microba pipeline means they can’t give you a good machine-readable breakdown of your results, which limits the appeal to serious microbiome hackers.
That said, with careful study I was able to identify oddities that I may not have found otherwise, so in a difficult-to-treat situation this may have made the difference in treatment. You certainly can’t beat the price.
Download my full report
See also May 2020 GutBiome+ Report