Viome Presentation at Personalized Nutrition Summit
Part 1 of my series of notes on presentations at the Personalized Nutrition Innovation Summit in June 2019

Momo Viyusich, Chief Science Officer at Viome, gave an impressive update on some of their latest scientific results. I hadn’t heard Momo’s personal story before, about his history of carpel tunnel syndrome and rheumatoid arthritis that was reversed when he switched to a “non-mammalian” diet. Apparently there is a dietary form of Sialic acid in mammalian cells that causes an autoimmune response in some people. When he dropped mammalian food from his diet, the symptoms went away, in contrast to the memory loss he suffered when he tried a ketogenic diet for three months.
Viome’s Business
A quick summary of their current business shows they are starting to accumulate a lot of data.
- 50K customers, growing now at 10K/month.
- 10 countries
- 170 employees (50 of whom are at their only lab, in Los Alamos)
Genes vs Nutrition
A fundamental belief at Viome is that genes alone have little or no predictive power for disease because it’s the expression of the genes that matters. You might have a gene that predisposes you to some condition, but if it’s never expressed, it doesn’t matter.
For example, one recent large study on genes associated with depression concluded:
hypotheses about depression candidate genes were incorrect and that the large number of associations reported in the depression candidate gene literature are likely to be false positives.1
Meanwhile, evidence continues to mount for the role of diet. For example the 2017 “SMILES” trial showed that depression is more effectively treated with dietary intervention than with social support therapy2
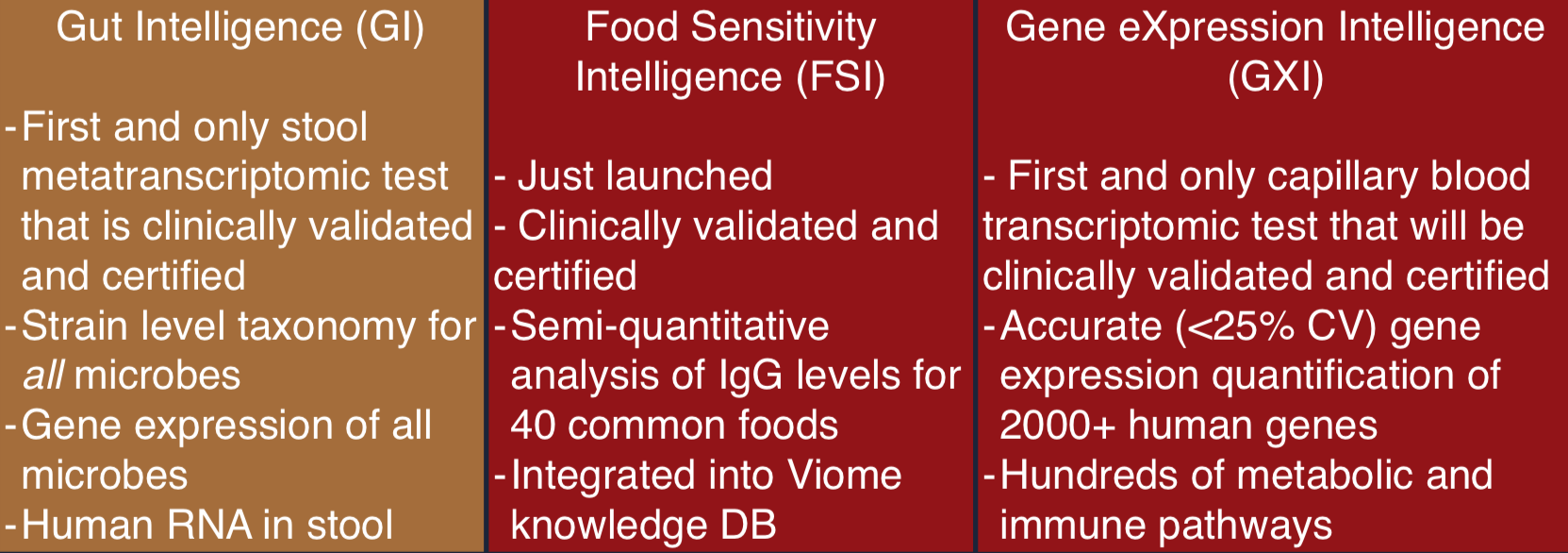
Viome Clinical Tests
The Viome pipeline is now 100% automated for existing tests, soon to be augmented with many others, including a new one for blood transcriptomics, and they may follow this eventually with other tests, including one for saliva.
Predictive Results
Even beyond the problems with trying to make associations based on genes alone, the vast majority of clinical studies are too underpowered to be useful. Unless you have a population of 50 or more (ideally hundreds), you can usually simply ignore the conclusions.
You can’t learn what actually matters from clinical studies under 50 (or even 100) people.
Here’s where Viome’s growing user database is coming into its own. In addition to the results published in May 20193 Momo presented some unpublished data showing some impressive predictive power of their algorithms on their large sample sizes. Using data from 1000 individuals, they claim to be able to predict conditions like obesity, IBS, type 2 diabetes and depression, with between 70 and 85% accuracy. They were also able to predict the rough age of the individual based purely on their microbiome results, speculating that it might be possible someday to “turn on” the microbial functions associated with youth.
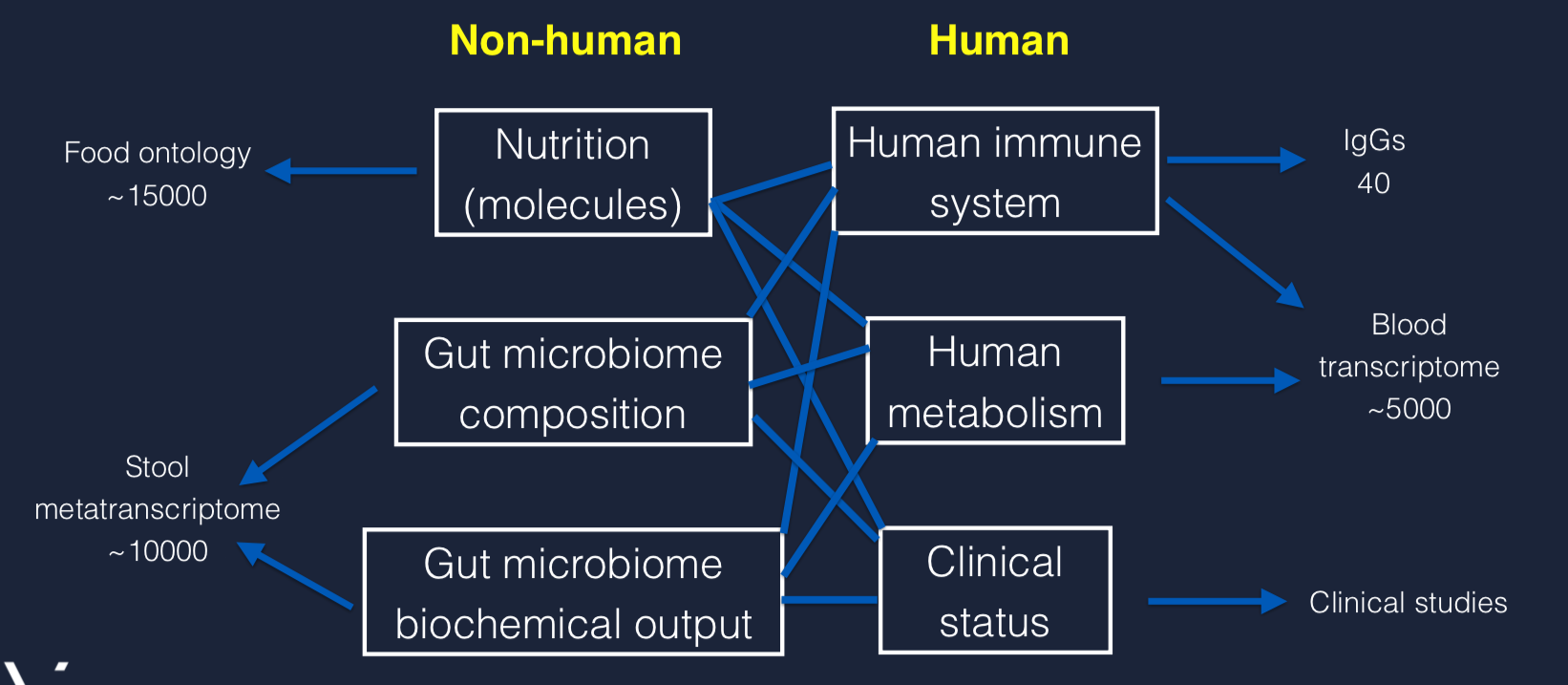
Some Technical Details
The Viome AI algorithm only looks at molecules, not food. They have a large database of foods and the molecules within. When building models of association between RNA expression and foods, they look only at the molecules. Food recommendations are later constructed in the reverse: foods that contain specific molecules.
Their “food ontology” consists of 15,000 molecules
Stool metatranscriptome: 10,000 sequences
Note that only about 10,000 of the 1M + genes of the microbiome are expressed.
Viome uses 2950 KEGG orthology
10M reads/sample of ribosomal RNA at 1:1M abundance sensitivity
Immune System: 40 IgGs
Blood Transcriptome: 50K sequences
The Future
Momo speculates that in five years they’d like to be able to tell the microbiome of somebody likely to develop pancreatic cancer, for example, in a way that would allow for an intervention to prevent it.
Meanwhile they want to focus more on practical ways to help people, like the personalized diet recommendations from Habit, the business they acquired in February 2019
For example, you could imagine an app that uses voice recognition to let you say something like “I have 30 minutes and would like some Thai food” — and it’ll make suggestions for specific menu items healthy for you.
Footnotes
Border, R., Johnson, E. C., Evans, L. M., Smolen, A., Berley, N., Sullivan, P. F., & Keller, M. C. (2019). No Support for Historical Candidate Gene or Candidate Gene-by-Interaction Hypotheses for Major Depression Across Multiple Large Samples. American Journal of Psychiatry, 176(5), 376–387. https://doi.org/10.1176/appi.ajp.2018.18070881]↩︎
Jacka, F. N., O’Neil, A., Opie, R., Itsiopoulos, C., Cotton, S., Mohebbi, M., … Berk, M. (2017). A randomised controlled trial of dietary improvement for adults with major depression (the ‘SMILES’ trial). BMC Medicine, 15(1). https://doi.org/10.1186/s12916-017-0791-y↩︎
Tily, H., Perlina, A., Patridge, E., Gline, S., Genkin, M., Gopu, V., … Banavar, G. (2019). Gut microbiome activity contributes to individual variation in glycemic response in adults. BioRxiv. https://doi.org/10.1101/641019↩︎